This web page was produced as an assignment for Genetics 564, an undergraduate course at UW‐Madison.
What are protein interaction networks?
Protein interaction networks are diagrams showing the interactions between a protein in interest and other proteins. These interactions can be direct (without intermediate) or indirect (with one or more intermediates).
Why study protein interaction networks?
Protein interaction networks illustrate the relationships between a protein of interest and its interacting partners, therefore provide an insight of the mechanisms or pathways of the biological functions of the protein of interest.
Methodology
The STRING database can be used to determine known and predicted protein interactions, which include direct, physical interactions and indirect associations inferred from genomic context, experiments, conserved coexpression, and data-mining in PubMed. [1]
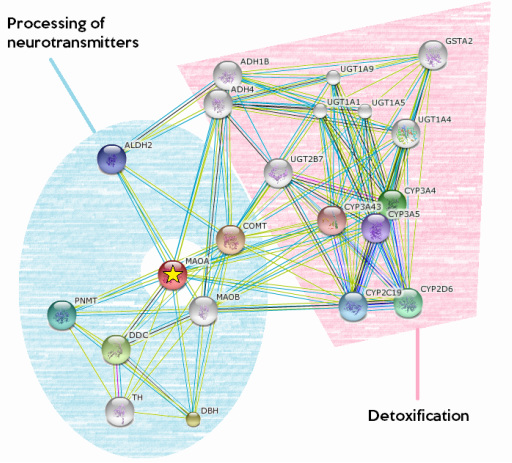
Protein interaction network of human MAOA
Proteins in the blue region are essential for the metabolism of neurotransmitter associated with emotional responses, such as serotonin, dopamin, epinephrine and norepinephrine.[2] Proteins in the pink region are mostly involved in detoxification, such as the break down of alcohol, testosterone, steroid, fatty acids and a number of therapeutic agents related to psychology and psychiatry. Given the fact that aggression is associated with a build-up of the mentioned neurotransmitters, testosterone and alcohol, the functions of the partners of MAOA protein are in conjunction with what I expected. A deeper look into the role of steroid and fatty acids in aggression and emotion processing would be interesting.
Reference:
- STRING - Known and Predicted Protein-Protein Interactions. Retrieved April 24, 2014.
- Buckholtz, J. W. & Meyer-Lindenberg, A. (2008). Maoa and the neurogenetic architecture of human aggression. Trends in Neurosciences, 31(3), 120-129.