This web page was produced as an assignment for Genetics 564, an undergraduate course at UW‐Madison.
What are homologous genes?
Homologous genes are hereditary units that derive from a common ancestor, consist of identical or similar sequences of nucleotides and control the same traits in different species. [1]
Why are gene homologs important?
Aggression is studied genetically based on its heritability through generations. Heritability models of aggression are mainly based on animals due to the ethical concern in using humans for genetic study. The presence of homologs allows scientists to study the genes in interest at much greater lengths using model organisms. Animals are first selectively bred, then placed in a variety of environmental conditions, allowing researchers to examine the differences in their expression of traits of aggression.
Homologs of MAOA gene
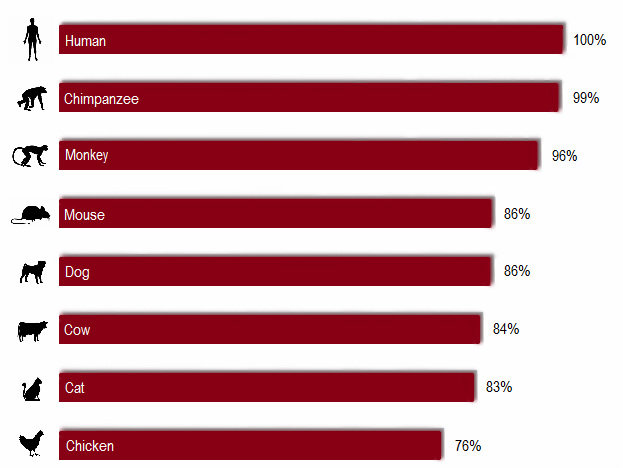
Figure 1. % identical of homologs in reference to the human MAOA gene
|
|
|
|
|
|
Chimpanzee is known to be human's closest evolutionary relative [2], and the analysis turned out as predicted: the most identical MAOA-encoding mRNA sequence to that of human's is found in chimpanzee.
Other than organisms listed above, homologs of human MAOA gene are also found in
Other than organisms listed above, homologs of human MAOA gene are also found in
- horse (equus caballus) - 86% identical
- brown rat (rattus norvegicus) - 86% identical
- sheep (ovis aries) - 84% identical
Methodology
The homologs of human MAOA gene mentioned above were identified using Homologene, an NCBI tool for detection of homologs among the annotated genes of several completely sequenced eukaryotic genomes. [3]
Next, sequence of each organism was compared to that of human's using BLAST2, another NCBI tool for detection of % identity between 2 sequences. Since it is impractical to compare genomic DNA sequences due to the presence of introns, the homology analysis was performed on mRNA sequences instead.
NOTE: BLAST2 result shows the two mRNA transcript variants of human MAOA are 100% identical. The isoform of human sequence used in the analysis for homology with other organisms was Transcript Variant 1.
Next, sequence of each organism was compared to that of human's using BLAST2, another NCBI tool for detection of % identity between 2 sequences. Since it is impractical to compare genomic DNA sequences due to the presence of introns, the homology analysis was performed on mRNA sequences instead.
NOTE: BLAST2 result shows the two mRNA transcript variants of human MAOA are 100% identical. The isoform of human sequence used in the analysis for homology with other organisms was Transcript Variant 1.
Discussion
Gene homology is useful for determining evolutionary history. The similarity of these homologs shows that MAOA is well conserved among mammals. However, gene homology is far less significant when it comes to determining similarity of protein structure and functions.
Note that sequence similarity might be due to convergent evolution (i.e. independent evolution of similar traits in different species) or merely because of chance, therefore homology can sometimes be incorrectly deduced on the basis of sequence similarity. [4] However, we can still identify partial homology by looking at the conserved regions of sequences under comparison. Regions of DNA sequences that share 100% identity in at least two different species are called "ultra-conserved elements" and have high functional value, since the probability of finding 100% identical regions by chance is extremely low. [5]
Note that sequence similarity might be due to convergent evolution (i.e. independent evolution of similar traits in different species) or merely because of chance, therefore homology can sometimes be incorrectly deduced on the basis of sequence similarity. [4] However, we can still identify partial homology by looking at the conserved regions of sequences under comparison. Regions of DNA sequences that share 100% identity in at least two different species are called "ultra-conserved elements" and have high functional value, since the probability of finding 100% identical regions by chance is extremely low. [5]
References:
- Homology. Merriam-Webster Online. Retrieved Feb 24, 2014.
- What are our closest animal relatives? Science Musuem. Retrived Feb 23, 2014.
- Homologene. National Center for Biotechnology Information. Retrieved Feb 23, 2014.
- Gauger, A. (2012) Similarity Happens! Biologic Institute. Retrieved March 21, 2014.
- Bejerano, G, Pheasant, M, Makunin, I, Stephen, S, Kent, WJ, Mattick, JS, & Haussler, D. (2004). Ultraconserved elements in the human genome. Science 304(5675): 1321–5.