This web page was produced as an assignment for Genetics 564, an undergraduate course at UW‐Madison.
What are homologous protein?
Homologous proteins are large biological molecules that derive from a common ancestor, consist of identical or similar sequences of nucleotides and perform the same functions in different species. [1] Homologous proteins have a very similar primary, secondary, and tertiary structure.
Why are protein homologs important?
In cases where human experimentation is ethically and financially inconvenient, the presence of homologs allows scientists to study the proteins in interest at much greater lengths using model organisms.
Homologs of MAOA proteins
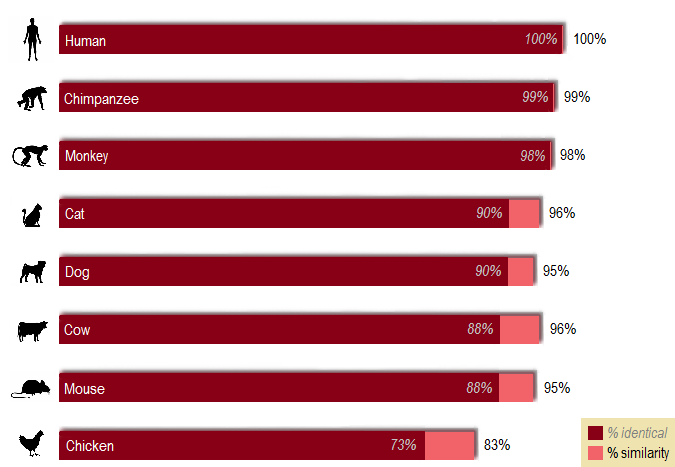
Figure 1. % identical and % similarity of homologs in reference to the human MAOA protein
|
|
|
|
|
|
|
|
Chimpanzee is known to be human's closest evolutionary relative [2], and the analysis turned out as predicted: the most identical MAOA protein sequence to that of human's is found in chimpanzee.
Other than organisms listed above, homologs of human MAOA protein are also found in
Other than organisms listed above, homologs of human MAOA protein are also found in
- horse (equus caballus) - 90% identical; 96% similar
- brown rat (rattus norvegicus) - 88% identical; 95% similar
- sheep (ovis aries) - 87% identical; 95% similar
% identity vs % similarity
% similarity of two sequences is either equal to or higher than their % identity, because % identity only calculates identical amino acids at specific positions, whereas % similarity measures the percentage of physicochemically equivalent amino acids (which can be identical or different amino acids). Different amino acids with equivalent physicochemical properties in theory would behave similarly, thus substitution of one by another usually does not cause dramatic changes to the overall structure or function of the protein. For example, leucine and valine are both hydrophobic and are expected to be equally tolerated within a hydrophobic core of a protein.
Methodology
The homologs of human MAOA protein mentioned above were identified using Homologene, an NCBI tool for detection of homologs among the annotated genes of several completely sequenced eukaryotic genomes. [3]
Next, sequence of each organism was compared to that of human's using BLAST2, another NCBI tool for detection of % identity and % similarity between 2 sequences.
NOTE: BLAST2 result shows the two isoforms of human MAOA protein are 100% identical. The isoform of human sequence used in the analysis for homology with other organisms was Variant 1.
Next, sequence of each organism was compared to that of human's using BLAST2, another NCBI tool for detection of % identity and % similarity between 2 sequences.
NOTE: BLAST2 result shows the two isoforms of human MAOA protein are 100% identical. The isoform of human sequence used in the analysis for homology with other organisms was Variant 1.
Discussion
Note that sequence similarity might be due to convergent evolution (i.e. independent evolution of similar traits in different species) or merely because of chance, therefore homology can sometimes be incorrectly deduced on the basis of sequence similarity. [4] However, we can still identify partial homology by looking at the conserved regions of sequences under comparison. The presence of identical or chemically equivalent amino acids implies conservation of protein sequences and protein structures. Highly conserved proteins are often essential for basic cellular functions.
References:
- Homology. Merriam-Webster Online. Retrieved Feb 24, 2014.
- What are our closest animal relatives? Science Musuem. Retrived Feb 23, 2014.
- Homologene. National Center for Biotechnology Information. Retrieved Feb 23, 2014.
- Gauger, A. (2012) Similarity Happens! Biologic Institute. Retrieved March 21, 2014.